
Align the cDNA sequence with the encoded protein sequence (using Expasy Translate tool). This gives the amino acid sequence as one letter codes with stop codons indicated by a hyphen.5. Select the sequence, copy it and then paste it into the translate sequence window in the ExPasy link. We have provided different protein sequencing using this tool below, so let’s check them out! 1. One such tool is an EXPASY Translate Tool which permits a nucleotide to create a protein sequence. There are different translation tools available that help a DNA translate its coding directly to proteins. In 25 years, SIB has grown from a handful of visionary scientists to a nationwide organization delivering essential services and resources to millions of users – researchers, clinicians, industry and governments alike.Transcription. A new identity to celebrate SIB’s 25 years and the future of life science data. Other resources that do not implement this specific interface (even if they have other query features), are not included in ExPASy’s parallel query.This ideo lecture describes How to use exapsy translate tool to translate DNA/RNA sequence to amino acid sequence/ protein sequenceSearch Queries: translate. If a resource implements the cross-resource search interface (B) (indicated by the red box named ‘Query interface’ and the arrow leading to the resource), ExPASy will query the resource directly. We would like to show you a description here but the site won’t allow us.translate in high-quality scientific databases and software tools using Expasy, the Swiss Bioinformatics Resource Portal. This resource is referenced here: Microbial Life:Research Methods:Genomics: Translation. 
If you have information we can use to flesh out or correct this record let us know. This description of a site outside SERC has not been vetted by SERC staff and may be incomplete or incorrect. Home Programmatic Access Contact Translate - Programmatic access Parameters dna_sequence Bare nucleotide sequence output_format The output format. Once the alignment is computed, you can view it using LALNVIEW, a graphical viewer program for pairwise alignments. 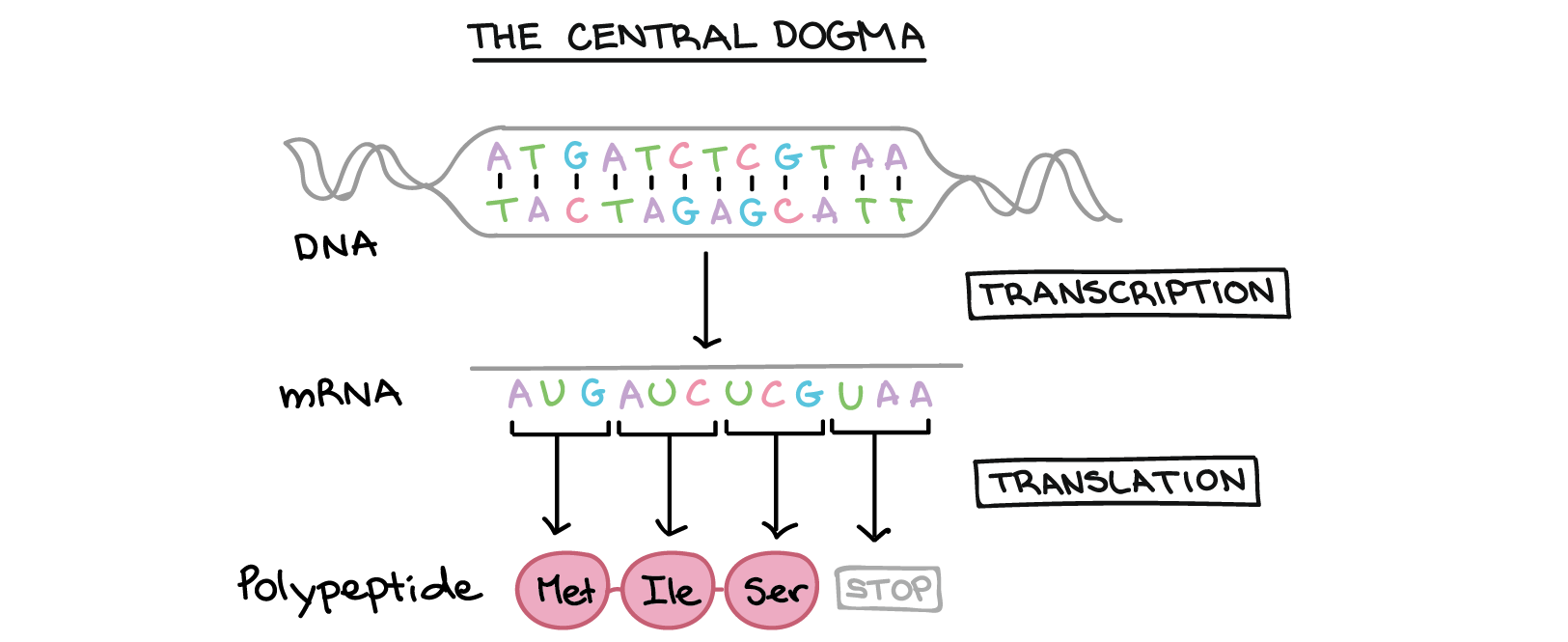
SIM is a program which finds a user-defined number of best non-intersecting alignments between two protein sequences or within a sequence. SIM - Alignment Tool for Protein Sequences. Try BLAST+ 2.14.1 today! Check out the changes we made. The program compares nucleotide or protein sequences to sequence databases and calculates the statistical significance. BLAST finds regions of similarity between biological sequences. I think you can and It may be possible by looking at the three frames (+) and choosing the one which is longest. Read our Privacy Notice if you are concerned with your privacy and how we handle personal information.If the sequences are not coding DNA sequences then stop codons are not relevant. If you plan to use these services during a course please contact us. If you have any feedback or encountered any issues please let us know via EMBL-EBI Support. 
Please read the provided Help & Documentation and FAQs before seeking help from our support staff. The tools described on this page are provided using Search and sequence analysis tools services from EMBL-EBI in 2022 Launch Sixpack Protein Sequence Back-translationĮMBOSS Backtranseq back-translates protein sequences to nucleotide sequences.ĮMBOSS Backtranambig back-translates protein sequences to ambiguous nucleotide sequences. Nucleotide Sequence TranslationĮMBOSS Transeq translates nucleic acid sequences to the corresponding peptide sequences.ĮMBOSS Sixpack displays DNA sequences with 6-frame translation and ORFs. Sequence Translation is used to translate nucleic acid sequence to corresponding peptide sequences.īack-translation is used to predict the possible nucleic acid sequence that a specified peptide sequence has originated from.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
